| TUD Organische Chemie | Immel | Tutorials | Orbitals | Molecular | Allyl-Cation, Anion, and Radical | View or Print (this frame only) |
The molecular orbitals (MOs) of molecules can be constructed by linear combination of atomic orbitals (LCAO).
Though the exact Schrödinger equation is unsolvable for many electron systems such
as molecules, the solution can be numerically approximated by ab initio or density functional (DFT) theory.
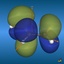
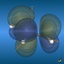
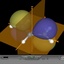
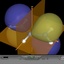
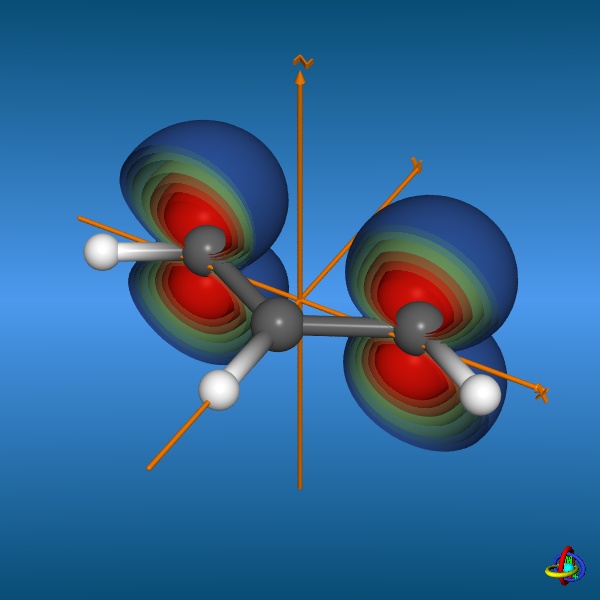
This page gives an overview on the molecular orbitals of the allyl-systems (cation, anion, and radical) calculated by DFT methods using a B3LYP/6-311++G** basis set. All MO representations are 90% or 90-25% iso-contour probability surfaces of the electron density (ψ2), i.e. they resemble the spatial volume around the nuclei of the molecule in which the electrons are found with the corresponding certainty. The different colors (yellow and blue) represent regions with opposite sign of the wave function ψ; nodal planes (not necessarily real "planar" planes) were ψ passes through zero and changes sign are indicated in orange.
Click on the small images below with blue background below to obtain an enlarged view - the images with black background provide links to the corresponding 3D-models (VRML-type models); these links will open in a new window. |
|
||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
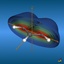
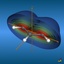
The total electron density (clipped 99, 95, 90, 80, 70, 60, and 50% iso-density contours depicted on the right) render the molecules with their characteristic shape
(note the different iso-contour values of the MO orbitals and the total electron density contours).
 |
|||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
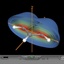
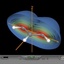
The total spin density (clipped 99, 95, 90, 80, 70, 60, and 50% iso-density contours depicted on the right) of the allyl-radical shows the the location
of the unpaired, single electron and its equal spreading over the two terminal carbon atoms.
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
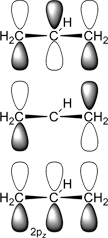
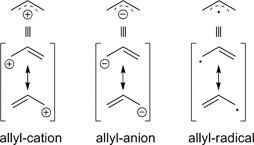
Below, plots for the molecular orbitals (only π-framework, σ-orbitals are not shown) of the allyl-cation and anion are given. The principal shapes of the corresponding
orbitals of the ally-cation and anion are very similar. As the allyl-cation has two electrons less than the anion, the lowest unoccupied molecular orbital (LUMO) of the cation
becomes the highest occupied molecular orbital (HOMO) in the anion (energies are not on scale in the diagrams below). For both species, the shapes of the frontier-orbitals govern their reactivities:
In the case of the cation, the LUMO of rather low energy interacts with nucleophiles (and the rather high-energy HOMO of the nucleophile), whereas the
high-energy HOMO of the anion interacts with electrophiles. In both cases (the LUMO of the cation and the HOMO of the anion) a nodal plane passes through the
central carbon atom, and only the terminal carbons contribute with AO coefficients not equal zero to these orbitals. Therefore, attack occurs invariably at the terminal positions only.
The thumbnail images below provide links to access to enlarged graphics as well as 3D-models (VRML) of the orbitals, respectively. Further informations on atomic orbitals (AOs) are available from the gallery of hydrogenic orbitals and hybrid orbitals. |
|||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
Notes: a) Nodal planes (ψ = 0.0); b) 90% Probability contours of MO electron density (ψ^2); c) 90, 80, 70, 60, 50, 40, and 25% Probability contours; d) 3D-Models require a VRML plugin to be installed (large files with sizes between 1000-5400 KBytes); files have been reduced in resolution to save band-width; e) Orbital not shown.
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||
For a more detailed description of atomic orbitals see the corresponding gallery of orbitals.
All graphics and iso-contour surfaces shown on this page were created using the MolArch+ program
and POVRAY Persistence of Vision Raytracer. Electron densities were calculated
on three dimensional grids for the corresponding molecules using the JAGUAR program.
|
|||||||||||||||||||||||||||||||||||||||||||||||||||||||||||||